New Genetic Engineering Toolset Produces Better-Modified Plants Faster, Easier

Researchers from NC State University have developed a set of tools that make it faster and easier to modify large segments of DNA.
The new tools are for use in a technique called recombineering. Recombineering is especially important for plant scientists, because many plants have particularly big genomes. A genome is the complete set of genetic material in a given organism. And some plant genomes can make a human genome seem puny.
For example, Paris japonica, a flowering plant from Japan, has a genome that is 50 times the size of a human’s. These huge genomes, and their correspondingly large individual genes, can make it challenging to introduce precise genetic changes to a plant. For example, making genetic modifications to help make a plant resistant to a new pest, or to study the fundamentals of how plants grow.
The researchers share how to use the new tools (and how they made them) in a paper published in the journal Plant Cell. They have also made the toolset available through the Arabidopsis Biological Resource Center.
“Before, unless you were in a specialized lab that worked with big pieces of DNA and had a lot of experience in molecular biology, it was unlikely you would adopt recombineering,” said Jose Alonso, a William Neal Reynolds professor of plant and microbial biology and senior author on the paper. “Our hope is that by making it more accessible, any molecular biology lab should be able to get into recombineering with these new tools. They should be a good resource.”
The toolset will improve the precision and reliability of fundamental plant science, he added. This basic science forms the foundation for innovations in pest resistance, yield increases and growing season extensions.
“With this toolset, we’re putting in an extra copy of the gene of interest — we’re not modifying the gene in the chromosome — but the gene is surrounded by the same large flanking DNA sequences that regulate it in the chromosome,” said Anna Stepanova, an associate professor of plant and microbial biology and an author on the paper. “This allows us to capture the normal environment of the gene and therefore when and where the gene is turned on and off.”
Including this contextual information allows plant scientists to gather more accurate information, whether they are interested in genes involved in plant growth or fighting off pests, she said.
The toolset can be used to introduce genetic modifications in almost any plant species, including crops like corn, rice and tomatoes, but it is easier to use in well-studied plants such as Arabidopsis, Alonso said. Arabidopsis is a small plant related to cabbage and mustard that serves as a “lab rat” for many plant scientists.
Genetic tools included in the toolset include those to add glowing tags to a protein of interest in order to track where it is made within growing plants, a tool to easily trim off excess or problematic DNA sequences and a tool to make it easier to transfer the newly modified gene of interest into a plant.
The toolset and the team’s protocols are scalable. They can be used to modify a single gene or used in high-throughput studies to investigate a whole genome, 96 genes at a time, Stepanova said. However, she expects most researchers will use the toolset to study 10 to 20 related proteins.
Researchers can also use the toolset to insert multiple modified genes into a single plant, even an entire series of genes to produce valuable compounds, such as certain medications, Stepanova said.
The toolset was built gradually over the past eight years as the tools were needed for other projects in the Alonso-Stepanova lab. Research scholars Javier Brumos and Chengsong Zhao were chiefly involved in creating the toolset, though three undergraduate students also assisted in the project.
“The students get familiarized with some of the molecular techniques and methodologies in recombineering, which is useful in whatever career they go into,” Alonso said. He added that one of the students, Yan Gong, is in graduate school at Stanford University; another, David Soriano, is interviewing at medical schools; and the third, Arjun Patel, is a junior at NC State.
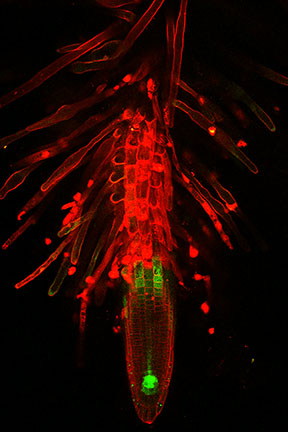
To test out the toolset, the research team added a yellow fluorescent tag to 96 genes involved in producing plant hormones. Then they transferred the tagged gene to bacteria that then inserted the tagged gene into the cells of Arabidopsis flowers. Some of the seedlings that grew from the seeds produced by these flowers received the modified gene and glowed yellow (under certain conditions). The researchers looked at when and where these glowing yellow proteins were produced in order to understand when and where plants make or respond to the growth hormone auxin.
The research was supported by a National Science Foundation grant to Alonso and seed funding from CALS Faculty Research and Professional Development program to Stepanova.
Our research addresses grand challenges — and overcomes them.
[button]Learn more[/button]
Note to Editors: The study abstract follows.
“An improved recombineering toolset for plants”
Authors: Javier Brumos, Chengsong Zhao, Yan Gong, David Soriano, Arjun P. Patel, Miguel A Perez-Amador, Anna N Stepanova and Jose M Alonso, North Carolina State University
Published: Oct. 30, The Plant Cell
DOI: 10.1105/tpc.19.00431
Abstract: Gene functional studies often rely on the expression of a gene of interest as transcriptional and translational fusions with specialized tags. Ideally, this is done in the native chromosomal contexts to avoid potential misexpression artifacts. Although recent improvements in genome editing make it possible to directly modify the target genes in their native chromosomal location, classical transgenesis is still the preferred experimental approach chosen in most gene tagging studies because of its time efficiency and accessibility. We have developed a recombineering-based tagging system that brings together the convenience of the classical transgenic approaches and the high degree of confidence in the obtained results provided by the direct chromosomal tagging achievable by genome editing strategies. These simple, scalable and customizable recombineering toolsets and protocols allow for the generation of a variety of genetic modifications. In addition, a highly efficient recombinase-mediated cassette exchange system has been developed to facilitate the transfer of the desired sequences from a BAC clone to a transformation-compatible binary vector, expanding the use of the recombineering approaches beyond Arabidopsis. The utility of this system is demonstrated by the generation of over 250 whole-gene translational fusions and 123 Arabidopsis transgenic lines corresponding to 62 auxin-related genes, and the characterization of the translational reporter expression patterns for 14 auxin biosynthesis genes.

